Tracks¶
Important
Since all tracks are hosted on the web with HTTP/HTTPS links provided for submission as tracks, the webservers which are hosting the track files need Cross-Origin Resource Sharing (CORS) enabled.
Quoted from MDN:
Cross-Origin Resource Sharing (CORS) is a mechanism that uses additional
HTTP headers to tell a browser to let a web application running at
one origin (domain) have permission to access selected resources
from a server at a different origin. A web application makes a
cross-origin HTTP request when it requests a resource that has
a different origin (domain, protocol, and port) than its own origin.
Configure your webserver to enable CORS¶
Most likely the browser domain is different from the server the tracks are hosted on. The hosting server needs CORS enabled and for an Apache web server in Ubuntu this setup will work:
Header always set Access-Control-Allow-Origin "*"
Header always set Access-Control-Allow-Methods "POST, GET, OPTIONS, DELETE, PUT"
Header always set Access-Control-Max-Age "1000"
Header always set Access-Control-Allow-Headers "x-requested-with, Content-Type, origin, authorization, accept, client-security-token"
Prepare track files¶
The browser accesses track files from their URL. Only a portion of the data, that within the specific view region, are transferred to the browser for visualization. Thus, all the track files need be hosted in a web accssible location using HTTP or HTTPS. The following sections introduce the track types that the browser supports.
Binary track file formats like bigWig and HiC can be used directly with the browser.
bedGraph, methylC, categorical, longrange and bed track files need to
be compressed by bgzip and indexed by tabix for use by the browser.
The resulting index file with suffix .tbi needs to be located
at the same URL with the .gz file.
Bed like format track files need be sorted before submission. For example, if we have a track file named track.bedgraph
we can use the generic Linux sort command, the bedSort tool from UCSC, or the sort-bed command from BEDOPS.
Here is an example command using each of the three methods:
# Using Linux sort
sort -k1,1 -k2,2n track.bedgraph > track.bedgraph.sorted
# Using bedSort
bedSort track.bedgraph track.bedgraph.sorted
# Using sort-bed
sort-bed track.bedgraph > track.bedgraph.sorted
Then the file must be compressed using bgzip and indexed using tabix:
bgzip track.bedgraph.sorted
tabix -p bed track.bedgraph.sorted.gz
Move files “track.bedgraph.sorted.gz” and “track.bedgraph.sorted.gz.tbi” to a web server. The two files must be in the same directory. Obtain the URL to “track.bedgraph.sorted.gz” for submission.
SAM files first need to be compressed to BAM files. BAM files need to be coordinate sorted and
indexed for use by the browser.
The resulting index file with suffix .bai needs be located
at the same URL with the .bam file.
Here is an example command:
# Using samtools view to convert to bam
samtools view -Sb test.sam > test.bam
# Using samtools sort to coordinate sort the file
samtools sort test.bam > test.sorted.bam
# Using samtools index
samtools index test.sorted.bam
Annotation Tracks¶
Annotation tracks represent genomic features or intervals across the genome. Popular examples include SNP files, CpG Island files, and blacklisted regions.
bed¶
bed format files can be used to annotate elements across the genome or to represent reads from a sequencing experiment.
For more about the bed format please check the UCSC bed page.
Example lines are below:
chr9 3035610 3036180 Blacklist_155 . +
chr9 3036200 3036480 Blacklist_156 . +
chr9 3036420 3036660 Blacklist_157 . +
Every line must consist of at least 3 fields separated by the Tab delimiter. The required fields from
left to right are chromosome, start position (0-based), and end position (not included).
A fourth (optional) column is reserved for the name of the interval and the sixth column (optional)
is reserved for the strand. All other columns are ignored, but can be present in the file.
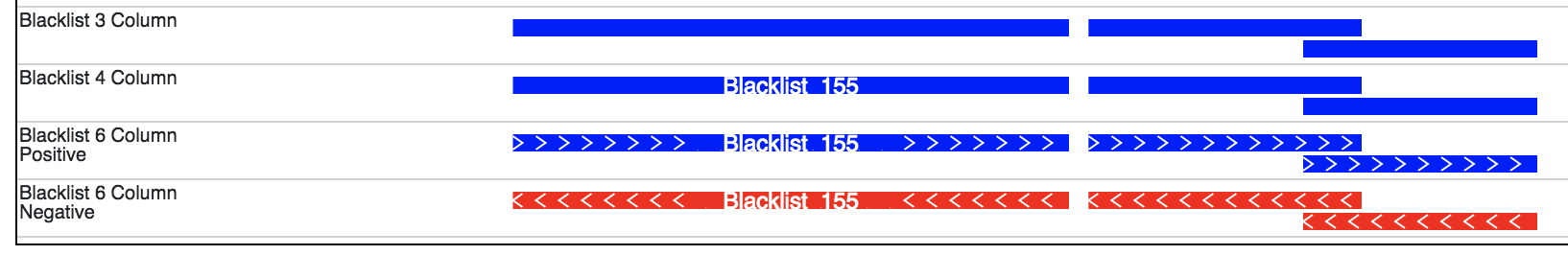
Note
The display of a bed file differs by how many columns are provided in the file
(see image above). The simplest, 3 column, format just displays blocks for
each interval. The four column format displays the name of each element over each interval.
If the sixth column is provided in the file then >>> or <<< will be displayed over
each interval to represent strand information.
This format needs to be compressed by bgzip and indexed by tabix for submission as a track. See Prepare track files.
refbed¶
The refbed format files allows you to upload a custom gene annotation track. It is similar to the
refGene bed-like file downloaded from UCSC but with slight modifications. Each file of
this format contains (each column is separated by Tab):
chr, transcript_start, transcript_stop, translation_start, translation_stop, strand, gene_name, transcript_id, type, exon(including UTR bases) starts, exon(including UTR bases) stops, and additional gene info (optional)
This format needs to be compressed by bgzip and indexed by tabix for submission as a track. See Prepare track files.
Hint
The 9th column contains gene type, but is simplified from the Gencode/Ensembl annotations to coding, pseudo, nonCoding, problem, and other. These classes of gene type are colored differently when the track is displayed on the browser.
Hint
The 10th and 11th columns contain exon starts and ends respectively. Each start or end is seperated by a comma.
For example:
start1,start2,start3,start4 stop1,stop2,stop3,stop4
100,120,140,160 110,130,150,170
Hint
The 12th column contains extra information. This information can be manually annotated or we suggest using Ensembl Biomart to download paired Transcript stable IDs and Gene descriptions. The information in this column must be seperated by spaces and not tabs.
All of the below lines will work for additional information in the 12th column:
Gene ID:ENSMUSG00000103482.1 Gene Type:TEC Transcript Type:TEC Additional Info:predicted gene, 37999 [Source:MGI Symbol;Acc:MGI:5611227]
Gene ID:ENSMUSG00000103482.1 Gene Type:TEC Transcript Type:TEC
ENSMUSG00000103482.1 TEC
Additional Info:predicted gene, 37999 [Source:MGI Symbol;Acc:MGI:5611227]
My Favorite Gene
Here are a few example lines in refbed format from gencode.vM17.annotation.gtf (mouse mm10 format):
chr1 24910461 24911659 24910461 24911659 - RP23-109H7.1 ENSMUST00000187022.1 pseudo 24911220,24910461 24911659,24910681 Gene ID:ENSMUSG00000100808.1 Gene Type:processed_pseudogene Transcript Type:processed_pseudogene Additional Info:predicted gene 28594 [Source:MGI Symbol;Acc:MGI:5579300]
chr1 25203443 25205696 25203443 25205696 - Adgrb3 ENSMUST00000190202.1 coding 25203443 25205696 Gene ID:ENSMUSG00000033569.17 Gene Type:protein_coding Transcript Type:retained_intron Additional Info:adhesion G protein-coupled receptor B3 [Source:MGI Symbol;Acc:MGI:2441837]
chr1 25276404 25277954 25276404 25277954 - RP23-21P2.4 ENSMUST00000193138.1 problem 25276404 25277954 Gene ID:ENSMUSG00000104257.1 Gene Type:TEC Transcript Type:TEC Additional Info:predicted gene, 20172 [Source:MGI Symbol;Acc:MGI:5012357]
chr1 26566833 26566938 26566833 26566938 + Gm24064 ENSMUST00000157486.1 nonCoding 26566833 26566938 Gene ID:ENSMUSG00000088111.1 Gene Type:snoRNA Transcript Type:snoRNA Additional Info:predicted gene, 24064 [Source:MGI Symbol;Acc:MGI:5453841]
Note
The last optional column is dislayed as a gene description when a gene is clicked on the browser. Our modified format can be
easily obtained from available refGene.bed file downloads from UCSC. Gencode GTF and Ensembl GTF files can be manipulated to
this format using the Converting_Gencode_or_Ensembl_GTF_to_refBed.bash script in scripts. The script by default puts
Gene ID:, Gene Type:, and Transcript Type: in the additional information column. Run with an annotation file, with
columns Transcript_ID and Description (seperated by a tab), the script will also add “Additional Info” to the 12th column. The
script depends on bedtools, bgzip, and tabix. Lastly, within the script an awk array is used to reclassify gene type and
can easily be modified for additional gene types.
The script is run as follows:
bash Converting_Gencode_or_Ensembl_GTF_to_refBed.bash Ensembl my.gtf my_optional_annotation.txt
bash Converting_Gencode_or_Ensembl_GTF_to_refBed.bash Gencode gencode.vM17.annotation.gtf
bash Converting_Gencode_or_Ensembl_GTF_to_refBed.bash Gencode gencode.vM17.annotation.gtf biomart_2col.txt
Warning
Spaces are used as delimiters in the GTF files so change gene names with spaces before processing.
For Example:
sed -i 's/ (1 of many)/_(1_of_many)/g' Danio_rerio.GRCz10.91.chr.gtf
Numerical Tracks¶
Currently there are two types of numerical tracks:
bigWig¶
bigWig is a popular format to represent numerical values over genomic coordinates.
Please check the UCSC bigWig page to learn more about this format.
bedGraph¶
bedGraph format also defines values in diffenent genomic locations.
For more about the bedGraph format please check the UCSC bedGraph page.
Example lines are below:
chr12 6537598 6537599 28.80914
chr12 6537599 6537600 28.96908
chr12 6537599 6537612 -2
chr12 6537600 6537601 29.30229
Every line consists of 4 fields separated by the Tab delimiter. The fields from
left to right are chromosome, start position (0-based), end position (not included), and value.
Note
You can use negative values for reverse strand. Both positive and negative
values can exist over the same coordinates (they can overlap). In bigWig format
negative values can also be specified, but they cannot overlap with positive values.
This format needs to be compressed by bgzip and indexed by tabix for submission as a track. See Prepare track files.
Read Alignment BAM Tracks¶
BAM¶
The BAM format is a compressed SAM format used to store sequence alignment data.
Please check the Samtools Documentation page to learn more about this format and how to manipulate these files.
Methylation tracks¶
Methylation experiments like MeDIP-seq or MRE-seq can use bigWig or bedGraph format for data display. For WGBS if users want to show read depth, methylation context, and methylation level then the data is best suited for the methylC format, described below.
methylC¶
Methylation data are formatted in methylC format, which is a 7 column bed format file:
chr1 10542 10543 CG 0.923 - 26
chr1 10556 10557 CHH 0.040 - 25
chr1 10562 10563 CG 0.941 + 17
chr1 10563 10564 CG 0.958 - 24
chr1 10564 10565 CHG 0.056 + 18
chr1 10566 10567 CHG 0.045 - 22
chr1 10570 10571 CG 0.870 + 23
chr1 10571 10572 CG 0.913 - 23
Each line contains 7 fields separated by Tab. The fields are
chromosome, start position (0-based), end position (not included),
methylation context (CG, CHG, CHG etc.), methylation value, strand,
and read depth.
This format needs to be compressed by bgzip and indexed by tabix for submission as a track. See Prepare track files.
Categorical tracks¶
Categorical tracks represent genomic bins for different categories. The most popular example is the represnetation of chromHMM data which indicates which region is likely an enhancer, likely a promoter, etc. Other uses for the track include the display of different types of methylation (DMRs, DMVs, LMRs, UMRs, etc.) or even peaks colored by tissue type.
categorical¶
The categorical track uses the first three columns of the standard bed format
(chromosome, start position (0-based), and end position (not included))
with the addition of a 4th column indicating the category type which can be a string or number:
chr1 start1 end1 category1
chr2 start2 end2 category2
chr3 start3 end3 category3
chr4 start4 end4 category4
Important
when you use numbers like 1, 2 and 3 as category names, in the datahub definition,
please use it a string for the category attribute in options, see the example below:
{
"type": "categorical",
"name": "ChromHMM",
"url": "https://egg.wustl.edu/d/hg19/E017_15_coreMarks_dense.gz",
"options": {
"category": {
"1": {"name": "Active TSS", "color": "#ff0000"},
"2": {"name": "Flanking Active TSS", "color": "#ff4500"},
"3": {"name": "Transcr at gene 5' and 3'", "color": "#32cd32"}
}
}
}
This format needs to be compressed by bgzip and indexed by tabix for submission as a track. See Prepare track files.
Long range chromatin interaction¶
Long range chromatin interaction data are used to show relationships between genomic regions. HiC is used to show the results from a HiC experiment.
HiC¶
To learn more about the HiC format please check https://github.com/aidenlab/juicer/wiki/Data.
longrange¶
The longrange track is a bed format-like file type. Each row contains columns from left to right:
chromosome, start position (0-based), and end position (not included), interaction target
in this format chr2:333-444,55. As an example, interval “chr1:111-222” interacts with
interval “chr2:333-444” on a score of 55,
we will use following two lines to represent this interaction:
chr1 111 222 chr2:333-444,55
chr2 333 444 chr1:111-222,55
Important
Be sure to make TWO records for a pair of interacting loci, one record for each locus.
This format needs to be compressed by bgzip and indexed by tabix for submission as a track. See Prepare track files.
bigInteract¶
The bigInteract format from UCSC can also be used at the browser, for more details about this format, please check the UCSC bigInteract format page.
cool¶
Thanks to the higlass team who provides the data API, the browser is able to display cool tracks by using the data uuid
from the higlass server, for example, you can use the uuid Hyc3TZevQVm3FcTAZShLQg to represent the track for Aiden et al. (2009) GM06900 HINDIII 1kb,
for a full list of available cool tracks please check http://higlass.io/api/v1/tilesets/?dt=matrix